Using older version of Lux.jl
This tutorial cannot be run on the latest Lux.jl release due to downstream packages not being updated yet.
Bayesian Neural Network
We borrow this tutorial from the official Turing Docs. We will show how the explicit parameterization of Lux enables first-class composability with packages which expect flattened out parameter vectors.
Note: The tutorial in the official Turing docs is now using Lux instead of Flux.
We will use Turing.jl with Lux.jl to implement implementing a classification algorithm. Lets start by importing the relevant libraries.
# Import libraries
using Lux, Turing, CairoMakie, Random, Tracker, Functors, LinearAlgebra
# Sampling progress
Turing.setprogress!(true);Precompiling Lux...
750.4 ms ✓ IteratorInterfaceExtensions
886.1 ms ✓ ConcreteStructs
793.3 ms ✓ CEnum
800.1 ms ✓ Future
808.5 ms ✓ ExprTools
817.1 ms ✓ OpenLibm_jll
825.2 ms ✓ ArgCheck
872.2 ms ✓ InverseFunctions
946.0 ms ✓ CompilerSupportLibraries_jll
1035.7 ms ✓ AbstractFFTs
1070.4 ms ✓ Statistics
1088.4 ms ✓ Serialization
1179.3 ms ✓ ADTypes
715.5 ms ✓ DataValueInterfaces
720.8 ms ✓ Reexport
1553.5 ms ✓ FillArrays
801.0 ms ✓ ManualMemory
716.3 ms ✓ SIMDTypes
797.0 ms ✓ RealDot
911.6 ms ✓ SuiteSparse_jll
918.1 ms ✓ OrderedCollections
917.6 ms ✓ Requires
1759.9 ms ✓ OffsetArrays
736.5 ms ✓ CompositionsBase
972.6 ms ✓ AbstractTrees
1955.3 ms ✓ UnsafeAtomics
909.8 ms ✓ IntervalSets
1010.8 ms ✓ EnzymeCore
728.1 ms ✓ IfElse
748.4 ms ✓ CommonWorldInvalidations
786.7 ms ✓ DataAPI
825.8 ms ✓ FastClosures
895.9 ms ✓ ConstructionBase
848.0 ms ✓ StaticArraysCore
1588.2 ms ✓ IrrationalConstants
1020.8 ms ✓ Compat
1133.4 ms ✓ CpuId
945.0 ms ✓ JLLWrappers
743.9 ms ✓ TableTraits
1150.0 ms ✓ DocStringExtensions
851.2 ms ✓ NaNMath
809.8 ms ✓ InverseFunctions → InverseFunctionsDatesExt
792.7 ms ✓ ChangesOfVariables → ChangesOfVariablesInverseFunctionsExt
869.2 ms ✓ RuntimeGeneratedFunctions
806.2 ms ✓ FillArrays → FillArraysStatisticsExt
755.9 ms ✓ CompositionsBase → CompositionsBaseInverseFunctionsExt
830.8 ms ✓ Adapt
1775.2 ms ✓ LazyArtifacts
747.6 ms ✓ IntervalSets → IntervalSetsRandomExt
752.7 ms ✓ IntervalSets → IntervalSetsStatisticsExt
759.4 ms ✓ ADTypes → ADTypesEnzymeCoreExt
956.4 ms ✓ Atomix
770.8 ms ✓ ConstructionBase → ConstructionBaseIntervalSetsExt
1504.6 ms ✓ ThreadingUtilities
781.1 ms ✓ ConstructionBase → ConstructionBaseLinearAlgebraExt
786.1 ms ✓ ADTypes → ADTypesConstructionBaseExt
2580.8 ms ✓ RecipesBase
844.0 ms ✓ DiffResults
778.7 ms ✓ Compat → CompatLinearAlgebraExt
1337.8 ms ✓ Static
1306.1 ms ✓ Hwloc_jll
1315.9 ms ✓ oneTBB_jll
3619.5 ms ✓ MacroTools
3009.9 ms ✓ Distributed
890.0 ms ✓ OffsetArrays → OffsetArraysAdaptExt
882.9 ms ✓ EnzymeCore → AdaptExt
1245.2 ms ✓ OpenSpecFun_jll
1412.7 ms ✓ FFTW_jll
1155.4 ms ✓ LogExpFunctions
1537.2 ms ✓ Tables
812.1 ms ✓ BitTwiddlingConvenienceFunctions
1035.5 ms ✓ IntervalSets → IntervalSetsRecipesBaseExt
859.4 ms ✓ LogExpFunctions → LogExpFunctionsInverseFunctionsExt
1203.2 ms ✓ CommonSubexpressions
2212.0 ms ✓ IntelOpenMP_jll
899.6 ms ✓ LogExpFunctions → LogExpFunctionsChangesOfVariablesExt
2498.8 ms ✓ LLVMExtra_jll
1967.2 ms ✓ ChainRulesCore
1806.3 ms ✓ CPUSummary
4940.8 ms ✓ Test
772.7 ms ✓ ADTypes → ADTypesChainRulesCoreExt
2523.9 ms ✓ DispatchDoctor
851.4 ms ✓ AbstractFFTs → AbstractFFTsChainRulesCoreExt
1156.2 ms ✓ PolyesterWeave
1083.3 ms ✓ InverseFunctions → InverseFunctionsTestExt
5671.3 ms ✓ SparseArrays
2948.9 ms ✓ IRTools
3252.3 ms ✓ Hwloc
2036.9 ms ✓ MKL_jll
853.9 ms ✓ DispatchDoctor → DispatchDoctorEnzymeCoreExt
1549.2 ms ✓ ChangesOfVariables → ChangesOfVariablesTestExt
1145.0 ms ✓ DispatchDoctor → DispatchDoctorChainRulesCoreExt
2006.1 ms ✓ LogExpFunctions → LogExpFunctionsChainRulesCoreExt
851.3 ms ✓ SuiteSparse
950.6 ms ✓ Statistics → SparseArraysExt
807.8 ms ✓ Hwloc → HwlocTrees
2174.6 ms ✓ AbstractFFTs → AbstractFFTsTestExt
1061.1 ms ✓ FillArrays → FillArraysSparseArraysExt
1152.6 ms ✓ ChainRulesCore → ChainRulesCoreSparseArraysExt
1005.2 ms ✓ SparseInverseSubset
1069.5 ms ✓ PDMats
1509.1 ms ✓ LuxCore
1441.3 ms ✓ Optimisers
847.8 ms ✓ FillArrays → FillArraysPDMatsExt
1684.0 ms ✓ ZygoteRules
745.6 ms ✓ LuxCore → LuxCoreEnzymeCoreExt
796.3 ms ✓ LuxCore → LuxCoreFunctorsExt
825.9 ms ✓ LuxCore → LuxCoreChainRulesCoreExt
675.6 ms ✓ Optimisers → OptimisersEnzymeCoreExt
2984.5 ms ✓ SpecialFunctions
8925.7 ms ✓ StaticArrays
782.9 ms ✓ StaticArrays → StaticArraysStatisticsExt
781.3 ms ✓ Adapt → AdaptStaticArraysExt
788.4 ms ✓ ConstructionBase → ConstructionBaseStaticArraysExt
813.0 ms ✓ StaticArrays → StaticArraysChainRulesCoreExt
735.3 ms ✓ GPUArraysCore
2129.7 ms ✓ SpecialFunctions → SpecialFunctionsChainRulesCoreExt
7322.7 ms ✓ LLVM
961.7 ms ✓ ArrayInterface
1099.2 ms ✓ StructArrays
1465.3 ms ✓ MLDataDevices
866.6 ms ✓ ArrayInterface → ArrayInterfaceChainRulesCoreExt
1072.2 ms ✓ ArrayInterface → ArrayInterfaceGPUArraysCoreExt
1136.7 ms ✓ ArrayInterface → ArrayInterfaceStaticArraysCoreExt
1339.6 ms ✓ DiffRules
1256.0 ms ✓ StructArrays → StructArraysLinearAlgebraExt
1357.5 ms ✓ StructArrays → StructArraysAdaptExt
1587.9 ms ✓ ArrayInterface → ArrayInterfaceSparseArraysExt
2633.3 ms ✓ Setfield
2104.6 ms ✓ StructArrays → StructArraysSparseArraysExt
1773.3 ms ✓ MLDataDevices → MLDataDevicesChainRulesCoreExt
1479.4 ms ✓ MLDataDevices → MLDataDevicesFillArraysExt
2329.1 ms ✓ StructArrays → StructArraysStaticArraysExt
1369.9 ms ✓ LuxCore → LuxCoreMLDataDevicesExt
1610.9 ms ✓ MLDataDevices → MLDataDevicesSparseArraysExt
1007.9 ms ✓ LuxCore → LuxCoreSetfieldExt
7660.0 ms ✓ FFTW
4470.4 ms ✓ Accessors
3571.8 ms ✓ UnsafeAtomicsLLVM
634.2 ms ✓ Accessors → AccessorsStructArraysExt
4385.5 ms ✓ WeightInitializers
861.2 ms ✓ Accessors → AccessorsStaticArraysExt
930.8 ms ✓ Accessors → AccessorsTestExt
965.9 ms ✓ Accessors → AccessorsIntervalSetsExt
968.1 ms ✓ Accessors → AccessorsDatesExt
1060.6 ms ✓ WeightInitializers → WeightInitializersChainRulesCoreExt
4395.1 ms ✓ ForwardDiff
990.1 ms ✓ ForwardDiff → ForwardDiffStaticArraysExt
4821.2 ms ✓ KernelAbstractions
21455.5 ms ✓ Unitful
768.0 ms ✓ Unitful → ConstructionBaseUnitfulExt
768.9 ms ✓ Unitful → InverseFunctionsUnitfulExt
828.6 ms ✓ Accessors → AccessorsUnitfulExt
1814.2 ms ✓ KernelAbstractions → LinearAlgebraExt
1841.3 ms ✓ KernelAbstractions → EnzymeExt
1849.6 ms ✓ StructArrays → StructArraysGPUArraysCoreExt
2078.5 ms ✓ KernelAbstractions → SparseArraysExt
4182.1 ms ✓ GPUArrays
5436.0 ms ✓ ChainRules
1812.1 ms ✓ MLDataDevices → MLDataDevicesGPUArraysExt
1922.6 ms ✓ WeightInitializers → WeightInitializersGPUArraysExt
954.4 ms ✓ ArrayInterface → ArrayInterfaceChainRulesExt
1050.7 ms ✓ MLDataDevices → MLDataDevicesChainRulesExt
6341.5 ms ✓ NNlib
1891.0 ms ✓ NNlib → NNlibForwardDiffExt
1894.4 ms ✓ NNlib → NNlibEnzymeCoreExt
1946.4 ms ✓ NNlib → NNlibFFTWExt
5558.5 ms ✓ Tracker
1986.9 ms ✓ ArrayInterface → ArrayInterfaceTrackerExt
2082.8 ms ✓ LuxCore → LuxCoreArrayInterfaceTrackerExt
2210.9 ms ✓ MLDataDevices → MLDataDevicesTrackerExt
2308.1 ms ✓ Tracker → TrackerPDMatsExt
1449.3 ms ✓ SymbolicIndexingInterface
1652.0 ms ✓ StaticArrayInterface
607.9 ms ✓ StaticArrayInterface → StaticArrayInterfaceOffsetArraysExt
798.0 ms ✓ StaticArrayInterface → StaticArrayInterfaceStaticArraysExt
614.0 ms ✓ CloseOpenIntervals
747.6 ms ✓ LayoutPointers
2228.9 ms ✓ RecursiveArrayTools
1119.7 ms ✓ StrideArraysCore
805.1 ms ✓ RecursiveArrayTools → RecursiveArrayToolsStructArraysExt
803.4 ms ✓ MLDataDevices → MLDataDevicesRecursiveArrayToolsExt
952.5 ms ✓ RecursiveArrayTools → RecursiveArrayToolsForwardDiffExt
1111.3 ms ✓ RecursiveArrayTools → RecursiveArrayToolsSparseArraysExt
888.1 ms ✓ Polyester
2082.5 ms ✓ RecursiveArrayTools → RecursiveArrayToolsTrackerExt
25812.1 ms ✓ Zygote
1788.5 ms ✓ MLDataDevices → MLDataDevicesZygoteExt
2783.2 ms ✓ Zygote → ZygoteTrackerExt
3172.7 ms ✓ RecursiveArrayTools → RecursiveArrayToolsZygoteExt
6469.4 ms ✓ LuxLib
4030.8 ms ✓ LuxLib → LuxLibTrackerExt
9768.4 ms ✓ Lux
2922.8 ms ✓ Lux → LuxTrackerExt
3700.0 ms ✓ Lux → LuxZygoteExt
195 dependencies successfully precompiled in 82 seconds. 31 already precompiled.
Precompiling Turing...
734.2 ms ✓ UnPack
850.0 ms ✓ LeftChildRightSiblingTrees
852.8 ms ✓ StackViews
886.9 ms ✓ Scratch
884.0 ms ✓ Showoff
921.1 ms ✓ DelimitedFiles
962.9 ms ✓ Missings
956.5 ms ✓ PaddedViews
1005.4 ms ✓ ProgressLogging
1010.4 ms ✓ LRUCache → SerializationExt
1025.1 ms ✓ LoggingExtras
1281.5 ms ✓ FunctionWrappers
1325.4 ms ✓ WoodburyMatrices
1345.2 ms ✓ SharedArrays
1548.6 ms ✓ ProgressMeter
778.6 ms ✓ SignedDistanceFields
1086.2 ms ✓ Libmount_jll
1061.7 ms ✓ LLVMOpenMP_jll
1090.1 ms ✓ Rmath_jll
1159.3 ms ✓ OpenSSL_jll
1097.6 ms ✓ Xorg_libXau_jll
1101.1 ms ✓ libpng_jll
1101.0 ms ✓ libfdk_aac_jll
1423.0 ms ✓ LogDensityProblems
1106.5 ms ✓ Imath_jll
1092.0 ms ✓ Giflib_jll
1098.9 ms ✓ LAME_jll
1811.9 ms ✓ SimpleTraits
1117.4 ms ✓ LERC_jll
1160.8 ms ✓ JpegTurbo_jll
1130.5 ms ✓ Xorg_libXdmcp_jll
1185.2 ms ✓ x265_jll
1250.6 ms ✓ XZ_jll
1160.3 ms ✓ Zstd_jll
1278.1 ms ✓ x264_jll
1200.3 ms ✓ Expat_jll
1288.0 ms ✓ libaom_jll
2597.6 ms ✓ UnicodeFun
1057.3 ms ✓ Xorg_xtrans_jll
1158.1 ms ✓ LZO_jll
1184.8 ms ✓ Opus_jll
1188.7 ms ✓ Libiconv_jll
1113.7 ms ✓ Libgpg_error_jll
3694.0 ms ✓ FixedPointNumbers
1060.7 ms ✓ Xorg_libpthread_stubs_jll
1132.2 ms ✓ FriBidi_jll
1081.2 ms ✓ Libuuid_jll
1202.5 ms ✓ DensityInterface
1087.5 ms ✓ Bzip2_jll
1126.4 ms ✓ Graphite2_jll
1106.4 ms ✓ EarCut_jll
1066.8 ms ✓ CRlibm_jll
1083.2 ms ✓ Ogg_jll
1049.8 ms ✓ isoband_jll
1084.3 ms ✓ Libffi_jll
1770.3 ms ✓ FilePathsBase
1120.1 ms ✓ L_BFGS_B_jll
1277.2 ms ✓ AxisArrays
796.0 ms ✓ SciMLStructures
924.2 ms ✓ Parameters
1442.2 ms ✓ DifferentiationInterface
1386.9 ms ✓ BangBang
2717.0 ms ✓ DataStructures
1180.4 ms ✓ FiniteDiff
3076.6 ms ✓ StringManipulation
850.2 ms ✓ RelocatableFolders
894.0 ms ✓ MosaicViews
1909.5 ms ✓ HypergeometricFunctions
2100.8 ms ✓ SplittablesBase
838.7 ms ✓ FunctionWrappersWrappers
931.5 ms ✓ ConsoleProgressMonitor
1220.5 ms ✓ TerminalLoggers
1258.1 ms ✓ Libtask
1313.1 ms ✓ AxisAlgorithms
1156.7 ms ✓ Pixman_jll
988.9 ms ✓ LogDensityProblemsAD
2737.3 ms ✓ SciMLOperators
1425.7 ms ✓ Rmath
1185.4 ms ✓ libsixel_jll
1313.2 ms ✓ OpenEXR_jll
1192.6 ms ✓ Libtiff_jll
829.1 ms ✓ Ratios → RatiosFixedPointNumbersExt
1255.9 ms ✓ Libgcrypt_jll
1313.7 ms ✓ XML2_jll
1307.6 ms ✓ FreeType2_jll
887.5 ms ✓ Isoband
1293.3 ms ✓ FilePathsBase → FilePathsBaseMmapExt
1134.9 ms ✓ LBFGSB
1637.3 ms ✓ libvorbis_jll
6178.2 ms ✓ PkgVersion
6143.8 ms ✓ FileIO
1054.8 ms ✓ DifferentiationInterface → DifferentiationInterfaceChainRulesCoreExt
5223.3 ms ✓ Roots
1591.7 ms ✓ DifferentiationInterface → DifferentiationInterfaceStaticArraysExt
2075.2 ms ✓ FilePathsBase → FilePathsBaseTestExt
1659.6 ms ✓ DifferentiationInterface → DifferentiationInterfaceForwardDiffExt
886.2 ms ✓ BangBang → BangBangChainRulesCoreExt
1299.3 ms ✓ DifferentiationInterface → DifferentiationInterfaceSparseArraysExt
1051.2 ms ✓ BangBang → BangBangStructArraysExt
986.6 ms ✓ BangBang → BangBangTablesExt
1421.7 ms ✓ BangBang → BangBangStaticArraysExt
6394.0 ms ✓ SparseConnectivityTracer
3819.2 ms ✓ ColorTypes
957.5 ms ✓ SortingAlgorithms
932.1 ms ✓ DifferentiationInterface → DifferentiationInterfaceFiniteDiffExt
1170.7 ms ✓ FiniteDiff → FiniteDiffSparseArraysExt
1238.4 ms ✓ FiniteDiff → FiniteDiffStaticArraysExt
4395.4 ms ✓ IntervalArithmetic
2048.6 ms ✓ SparseMatrixColorings
2717.2 ms ✓ DifferentiationInterface → DifferentiationInterfaceZygoteExt
999.2 ms ✓ LogDensityProblemsAD → LogDensityProblemsADADTypesExt
3525.0 ms ✓ DifferentiationInterface → DifferentiationInterfaceTrackerExt
1968.8 ms ✓ NamedArrays
954.4 ms ✓ LogDensityProblemsAD → LogDensityProblemsADDifferentiationInterfaceExt
1740.1 ms ✓ QuadGK
1383.1 ms ✓ LogDensityProblemsAD → LogDensityProblemsADForwardDiffExt
981.9 ms ✓ SciMLOperators → SciMLOperatorsStaticArraysCoreExt
971.4 ms ✓ Roots → RootsChainRulesCoreExt
1302.9 ms ✓ Gettext_jll
1311.2 ms ✓ XSLT_jll
1446.6 ms ✓ SciMLOperators → SciMLOperatorsSparseArraysExt
1475.6 ms ✓ Fontconfig_jll
1364.9 ms ✓ Roots → RootsForwardDiffExt
2757.2 ms ✓ LogDensityProblemsAD → LogDensityProblemsADZygoteExt
1515.9 ms ✓ FilePaths
2089.6 ms ✓ FreeType
1658.2 ms ✓ MicroCollections
11489.5 ms ✓ SIMD
3360.2 ms ✓ LogDensityProblemsAD → LogDensityProblemsADTrackerExt
3160.1 ms ✓ StatsFuns
3438.9 ms ✓ Interpolations
2662.6 ms ✓ SparseConnectivityTracer → SparseConnectivityTracerNaNMathExt
1580.3 ms ✓ IntervalArithmetic → IntervalArithmeticForwardDiffExt
1216.9 ms ✓ IntervalArithmetic → IntervalArithmeticIntervalSetsExt
2834.3 ms ✓ SparseConnectivityTracer → SparseConnectivityTracerLogExpFunctionsExt
2148.6 ms ✓ NLSolversBase
1247.7 ms ✓ IntervalArithmetic → IntervalArithmeticDiffRulesExt
8906.6 ms ✓ GeometryBasics
1444.0 ms ✓ IntervalArithmetic → IntervalArithmeticRecipesBaseExt
3100.1 ms ✓ SparseConnectivityTracer → SparseConnectivityTracerSpecialFunctionsExt
3062.6 ms ✓ QOI
1792.5 ms ✓ DifferentiationInterface → DifferentiationInterfaceSparseMatrixColoringsExt
3733.9 ms ✓ ColorVectorSpace
1728.8 ms ✓ Glib_jll
1221.4 ms ✓ StatsFuns → StatsFunsInverseFunctionsExt
4380.3 ms ✓ SparseConnectivityTracer → SparseConnectivityTracerNNlibExt
3859.3 ms ✓ StatsBase
2163.0 ms ✓ Xorg_libxcb_jll
1981.9 ms ✓ Interpolations → InterpolationsUnitfulExt
2688.0 ms ✓ StatsFuns → StatsFunsChainRulesCoreExt
1823.7 ms ✓ Packing
1272.5 ms ✓ ColorVectorSpace → SpecialFunctionsExt
1194.4 ms ✓ Xorg_libX11_jll
3097.4 ms ✓ LineSearches
2324.5 ms ✓ ShaderAbstractions
6315.9 ms ✓ Colors
4112.8 ms ✓ Transducers
1168.8 ms ✓ Xorg_libXext_jll
1186.8 ms ✓ Xorg_libXrender_jll
1079.5 ms ✓ Animations
2124.6 ms ✓ ColorBrewer
2735.6 ms ✓ OpenEXR
1700.8 ms ✓ SparseMatrixColorings → SparseMatrixColoringsColorsExt
5150.3 ms ✓ ExactPredicates
3405.4 ms ✓ FreeTypeAbstraction
1588.1 ms ✓ Transducers → TransducersAdaptExt
3336.5 ms ✓ Zygote → ZygoteColorsExt
6204.8 ms ✓ MakieCore
1395.7 ms ✓ Cairo_jll
1508.6 ms ✓ Libglvnd_jll
4966.4 ms ✓ Optim
1407.5 ms ✓ HarfBuzz_jll
1425.2 ms ✓ libwebp_jll
8723.3 ms ✓ GridLayoutBase
5547.6 ms ✓ ColorSchemes
2664.2 ms ✓ AbstractMCMC
1107.2 ms ✓ libass_jll
7718.9 ms ✓ Distributions
1618.0 ms ✓ FFMPEG_jll
1954.1 ms ✓ SSMProblems
2368.3 ms ✓ AbstractPPL
2078.3 ms ✓ Distributions → DistributionsDensityInterfaceExt
2197.0 ms ✓ Distributions → DistributionsChainRulesCoreExt
2386.5 ms ✓ Distributions → DistributionsTestExt
7402.2 ms ✓ DelaunayTriangulation
14458.5 ms ✓ Automa
4458.3 ms ✓ AdvancedHMC
2428.5 ms ✓ MCMCDiagnosticTools
2628.5 ms ✓ EllipticalSliceSampling
3270.9 ms ✓ KernelDensity
3216.0 ms ✓ AdvancedMH
3289.0 ms ✓ AdvancedPS
17590.1 ms ✓ SciMLBase
5873.3 ms ✓ Bijectors
2819.9 ms ✓ AdvancedMH → AdvancedMHStructArraysExt
2912.1 ms ✓ AdvancedMH → AdvancedMHForwardDiffExt
24736.7 ms ✓ PrettyTables
1983.5 ms ✓ SciMLBase → SciMLBaseChainRulesCoreExt
6553.8 ms ✓ DistributionsAD
3630.6 ms ✓ AdvancedPS → AdvancedPSLibtaskExt
2330.1 ms ✓ Bijectors → BijectorsForwardDiffExt
2534.9 ms ✓ DistributionsAD → DistributionsADForwardDiffExt
2721.2 ms ✓ Bijectors → BijectorsDistributionsADExt
5239.7 ms ✓ SciMLBase → SciMLBaseZygoteExt
13047.1 ms ✓ PlotUtils
4703.6 ms ✓ Bijectors → BijectorsZygoteExt
4393.9 ms ✓ MCMCChains
5204.4 ms ✓ Bijectors → BijectorsTrackerExt
4595.8 ms ✓ DistributionsAD → DistributionsADTrackerExt
3030.6 ms ✓ AdvancedMH → AdvancedMHMCMCChainsExt
3077.3 ms ✓ AdvancedHMC → AdvancedHMCMCMCChainsExt
23313.7 ms ✓ ImageCore
3616.2 ms ✓ AdvancedVI
2813.7 ms ✓ ImageBase
16226.2 ms ✓ MathTeXEngine
4074.6 ms ✓ WebP
4441.9 ms ✓ PNGFiles
4139.1 ms ✓ AdvancedVI → AdvancedVIZygoteExt
4883.7 ms ✓ JpegTurbo
5069.5 ms ✓ Sixel
2431.2 ms ✓ ImageAxes
1385.1 ms ✓ ImageMetadata
10374.5 ms ✓ DynamicPPL
2337.0 ms ✓ Netpbm
2094.7 ms ✓ DynamicPPL → DynamicPPLEnzymeCoreExt
2149.6 ms ✓ DynamicPPL → DynamicPPLForwardDiffExt
2260.2 ms ✓ DynamicPPL → DynamicPPLChainRulesCoreExt
2809.1 ms ✓ DynamicPPL → DynamicPPLMCMCChainsExt
3089.9 ms ✓ DynamicPPL → DynamicPPLZygoteRulesExt
57410.7 ms ✓ TiffImages
1422.1 ms ✓ ImageIO
112785.7 ms ✓ Makie
9018.3 ms ✓ SciMLBase → SciMLBaseMakieExt
2241.1 ms ✓ OptimizationBase
513.2 ms ✓ OptimizationBase → OptimizationFiniteDiffExt
796.3 ms ✓ OptimizationBase → OptimizationForwardDiffExt
1541.4 ms ✓ OptimizationBase → OptimizationMLDataDevicesExt
2339.1 ms ✓ OptimizationBase → OptimizationZygoteExt
2085.7 ms ✓ Optimization
12195.8 ms ✓ OptimizationOptimJL
5958.3 ms ✓ Turing
4926.6 ms ✓ Turing → TuringOptimExt
242 dependencies successfully precompiled in 227 seconds. 237 already precompiled.
Precompiling CairoMakie...
711.8 ms ✓ Graphics
926.8 ms ✓ Pango_jll
1408.3 ms ✓ Cairo
74662.6 ms ✓ CairoMakie
4 dependencies successfully precompiled in 77 seconds. 292 already precompiled.
[ Info: [Turing]: progress logging is enabled globally
[ Info: [AdvancedVI]: global PROGRESS is set as trueGenerating data
Our goal here is to use a Bayesian neural network to classify points in an artificial dataset. The code below generates data points arranged in a box-like pattern and displays a graph of the dataset we'll be working with.
# Number of points to generate
N = 80
M = round(Int, N / 4)
rng = Random.default_rng()
Random.seed!(rng, 1234)
# Generate artificial data
x1s = rand(rng, Float32, M) * 4.5f0;
x2s = rand(rng, Float32, M) * 4.5f0;
xt1s = Array([[x1s[i] + 0.5f0; x2s[i] + 0.5f0] for i in 1:M])
x1s = rand(rng, Float32, M) * 4.5f0;
x2s = rand(rng, Float32, M) * 4.5f0;
append!(xt1s, Array([[x1s[i] - 5.0f0; x2s[i] - 5.0f0] for i in 1:M]))
x1s = rand(rng, Float32, M) * 4.5f0;
x2s = rand(rng, Float32, M) * 4.5f0;
xt0s = Array([[x1s[i] + 0.5f0; x2s[i] - 5.0f0] for i in 1:M])
x1s = rand(rng, Float32, M) * 4.5f0;
x2s = rand(rng, Float32, M) * 4.5f0;
append!(xt0s, Array([[x1s[i] - 5.0f0; x2s[i] + 0.5f0] for i in 1:M]))
# Store all the data for later
xs = [xt1s; xt0s]
ts = [ones(2 * M); zeros(2 * M)]
# Plot data points
function plot_data()
x1 = first.(xt1s)
y1 = last.(xt1s)
x2 = first.(xt0s)
y2 = last.(xt0s)
fig = Figure()
ax = CairoMakie.Axis(fig[1, 1]; xlabel="x", ylabel="y")
scatter!(ax, x1, y1; markersize=16, color=:red, strokecolor=:black, strokewidth=2)
scatter!(ax, x2, y2; markersize=16, color=:blue, strokecolor=:black, strokewidth=2)
return fig
end
plot_data()Building the Neural Network
The next step is to define a feedforward neural network where we express our parameters as distributions, and not single points as with traditional neural networks. For this we will use Dense to define liner layers and compose them via Chain, both are neural network primitives from Lux. The network nn we will create will have two hidden layers with tanh activations and one output layer with sigmoid activation, as shown below.
The nn is an instance that acts as a function and can take data, parameters and current state as inputs and output predictions. We will define distributions on the neural network parameters.
# Construct a neural network using Lux
nn = Chain(Dense(2 => 3, tanh), Dense(3 => 2, tanh), Dense(2 => 1, sigmoid))
# Initialize the model weights and state
ps, st = Lux.setup(rng, nn)
Lux.parameterlength(nn) # number of parameters in NN20The probabilistic model specification below creates a parameters variable, which has IID normal variables. The parameters represents all parameters of our neural net (weights and biases).
# Create a regularization term and a Gaussian prior variance term.
alpha = 0.09
sig = sqrt(1.0 / alpha)3.3333333333333335Construct named tuple from a sampled parameter vector. We could also use ComponentArrays here and simply broadcast to avoid doing this. But let's do it this way to avoid dependencies.
function vector_to_parameters(ps_new::AbstractVector, ps::NamedTuple)
@assert length(ps_new) == Lux.parameterlength(ps)
i = 1
function get_ps(x)
z = reshape(view(ps_new, i:(i + length(x) - 1)), size(x))
i += length(x)
return z
end
return fmap(get_ps, ps)
endvector_to_parameters (generic function with 1 method)To interface with external libraries it is often desirable to use the StatefulLuxLayer to automatically handle the neural network states.
const model = StatefulLuxLayer{true}(nn, nothing, st)
# Specify the probabilistic model.
@model function bayes_nn(xs, ts)
# Sample the parameters
nparameters = Lux.parameterlength(nn)
parameters ~ MvNormal(zeros(nparameters), Diagonal(abs2.(sig .* ones(nparameters))))
# Forward NN to make predictions
preds = Lux.apply(model, xs, vector_to_parameters(parameters, ps))
# Observe each prediction.
for i in eachindex(ts)
ts[i] ~ Bernoulli(preds[i])
end
endbayes_nn (generic function with 2 methods)Inference can now be performed by calling sample. We use the HMC sampler here.
# Perform inference.
N = 5000
ch = sample(bayes_nn(reduce(hcat, xs), ts), HMC(0.05, 4; adtype=AutoTracker()), N)Chains MCMC chain (5000×30×1 Array{Float64, 3}):
Iterations = 1:1:5000
Number of chains = 1
Samples per chain = 5000
Wall duration = 22.87 seconds
Compute duration = 22.87 seconds
parameters = parameters[1], parameters[2], parameters[3], parameters[4], parameters[5], parameters[6], parameters[7], parameters[8], parameters[9], parameters[10], parameters[11], parameters[12], parameters[13], parameters[14], parameters[15], parameters[16], parameters[17], parameters[18], parameters[19], parameters[20]
internals = lp, n_steps, is_accept, acceptance_rate, log_density, hamiltonian_energy, hamiltonian_energy_error, numerical_error, step_size, nom_step_size
Summary Statistics
parameters mean std mcse ess_bulk ess_tail rhat ess_per_sec
Symbol Float64 Float64 Float64 Float64 Float64 Float64 Float64
parameters[1] 5.8536 2.5579 0.6449 16.6210 21.2567 1.2169 0.7268
parameters[2] 0.1106 0.3642 0.0467 76.1493 35.8893 1.0235 3.3300
parameters[3] 4.1685 2.2970 0.6025 15.9725 62.4537 1.0480 0.6985
parameters[4] 1.0580 1.9179 0.4441 22.3066 51.3818 1.0513 0.9755
parameters[5] 4.7925 2.0622 0.5484 15.4001 28.2539 1.1175 0.6734
parameters[6] 0.7155 1.3734 0.2603 28.7492 59.2257 1.0269 1.2572
parameters[7] 0.4981 2.7530 0.7495 14.5593 22.0260 1.2506 0.6367
parameters[8] 0.4568 1.1324 0.2031 31.9424 38.7102 1.0447 1.3968
parameters[9] -1.0215 2.6186 0.7268 14.2896 22.8493 1.2278 0.6249
parameters[10] 2.1324 1.6319 0.4231 15.0454 43.2111 1.3708 0.6579
parameters[11] -2.0262 1.8130 0.4727 15.0003 23.5212 1.2630 0.6560
parameters[12] -4.5525 1.9168 0.4399 18.6812 29.9668 1.0581 0.8169
parameters[13] 3.7207 1.3736 0.2889 22.9673 55.7445 1.0128 1.0043
parameters[14] 2.5799 1.7626 0.4405 17.7089 38.8364 1.1358 0.7744
parameters[15] -1.3181 1.9554 0.5213 14.6312 22.0160 1.1793 0.6398
parameters[16] -2.9322 1.2308 0.2334 28.3970 130.8667 1.0216 1.2418
parameters[17] -2.4957 2.7976 0.7745 16.2068 20.1562 1.0692 0.7087
parameters[18] -5.0880 1.1401 0.1828 39.8971 52.4786 1.1085 1.7447
parameters[19] -4.7674 2.0627 0.5354 21.4562 18.3886 1.0764 0.9383
parameters[20] -4.7466 1.2214 0.2043 38.5170 32.7162 1.0004 1.6843
Quantiles
parameters 2.5% 25.0% 50.0% 75.0% 97.5%
Symbol Float64 Float64 Float64 Float64 Float64
parameters[1] 0.9164 4.2536 5.9940 7.2512 12.0283
parameters[2] -0.5080 -0.1044 0.0855 0.2984 1.0043
parameters[3] 0.3276 2.1438 4.2390 6.1737 7.8532
parameters[4] -1.4579 -0.1269 0.4550 1.6893 5.8331
parameters[5] 1.4611 3.3711 4.4965 5.6720 9.3282
parameters[6] -1.2114 -0.1218 0.4172 1.2724 4.1938
parameters[7] -6.0297 -0.5712 0.5929 2.1686 5.8786
parameters[8] -1.8791 -0.2492 0.4862 1.1814 2.9032
parameters[9] -6.7656 -2.6609 -0.4230 0.9269 2.8021
parameters[10] -1.2108 1.0782 2.0899 3.3048 5.0428
parameters[11] -6.1454 -3.0731 -2.0592 -1.0526 1.8166
parameters[12] -8.8873 -5.8079 -4.2395 -3.2409 -1.2353
parameters[13] 1.2909 2.6693 3.7502 4.6268 6.7316
parameters[14] -0.2741 1.2807 2.2801 3.5679 6.4876
parameters[15] -4.7115 -2.6584 -1.4956 -0.2644 3.3498
parameters[16] -5.4427 -3.7860 -2.8946 -1.9382 -0.8417
parameters[17] -6.4221 -4.0549 -2.9178 -1.7934 5.5835
parameters[18] -7.5413 -5.8069 -5.0388 -4.3025 -3.0121
parameters[19] -7.2611 -5.9449 -5.2768 -4.3663 2.1958
parameters[20] -7.0130 -5.5204 -4.8727 -3.9813 -1.9280Now we extract the parameter samples from the sampled chain as θ (this is of size 5000 x 20 where 5000 is the number of iterations and 20 is the number of parameters). We'll use these primarily to determine how good our model's classifier is.
# Extract all weight and bias parameters.
θ = MCMCChains.group(ch, :parameters).value;Prediction Visualization
# A helper to run the nn through data `x` using parameters `θ`
nn_forward(x, θ) = model(x, vector_to_parameters(θ, ps))
# Plot the data we have.
fig = plot_data()
# Find the index that provided the highest log posterior in the chain.
_, i = findmax(ch[:lp])
# Extract the max row value from i.
i = i.I[1]
# Plot the posterior distribution with a contour plot
x1_range = collect(range(-6; stop=6, length=25))
x2_range = collect(range(-6; stop=6, length=25))
Z = [nn_forward([x1, x2], θ[i, :])[1] for x1 in x1_range, x2 in x2_range]
contour!(x1_range, x2_range, Z; linewidth=3, colormap=:seaborn_bright)
figThe contour plot above shows that the MAP method is not too bad at classifying our data. Now we can visualize our predictions.
The nn_predict function takes the average predicted value from a network parameterized by weights drawn from the MCMC chain.
# Return the average predicted value across multiple weights.
nn_predict(x, θ, num) = mean([first(nn_forward(x, view(θ, i, :))) for i in 1:10:num])nn_predict (generic function with 1 method)Next, we use the nn_predict function to predict the value at a sample of points where the x1 and x2 coordinates range between -6 and 6. As we can see below, we still have a satisfactory fit to our data, and more importantly, we can also see where the neural network is uncertain about its predictions much easier–-those regions between cluster boundaries.
Plot the average prediction.
fig = plot_data()
n_end = 1500
x1_range = collect(range(-6; stop=6, length=25))
x2_range = collect(range(-6; stop=6, length=25))
Z = [nn_predict([x1, x2], θ, n_end)[1] for x1 in x1_range, x2 in x2_range]
contour!(x1_range, x2_range, Z; linewidth=3, colormap=:seaborn_bright)
figSuppose we are interested in how the predictive power of our Bayesian neural network evolved between samples. In that case, the following graph displays an animation of the contour plot generated from the network weights in samples 1 to 5,000.
fig = plot_data()
Z = [first(nn_forward([x1, x2], θ[1, :])) for x1 in x1_range, x2 in x2_range]
c = contour!(x1_range, x2_range, Z; linewidth=3, colormap=:seaborn_bright)
record(fig, "results.gif", 1:250:size(θ, 1)) do i
fig.current_axis[].title = "Iteration: $i"
Z = [first(nn_forward([x1, x2], θ[i, :])) for x1 in x1_range, x2 in x2_range]
c[3] = Z
return fig
end"results.gif"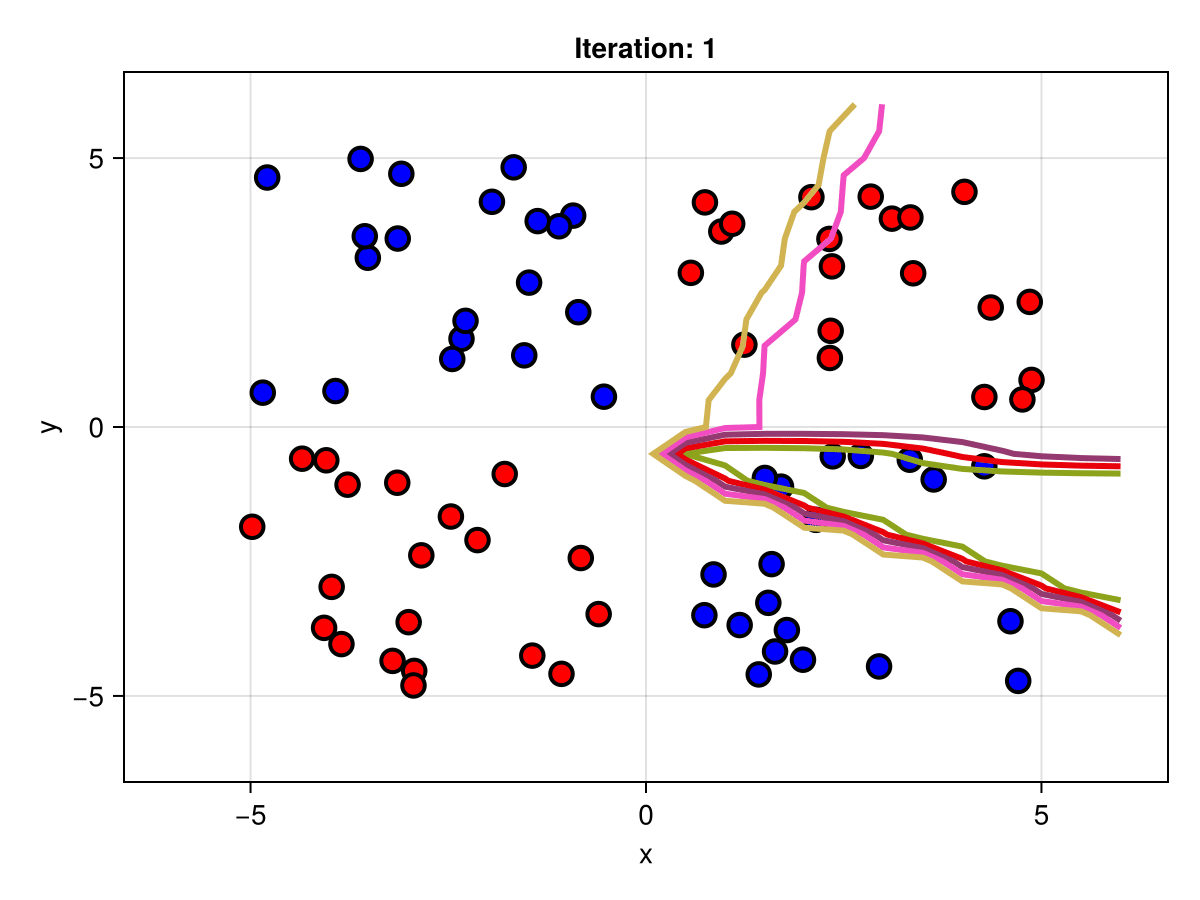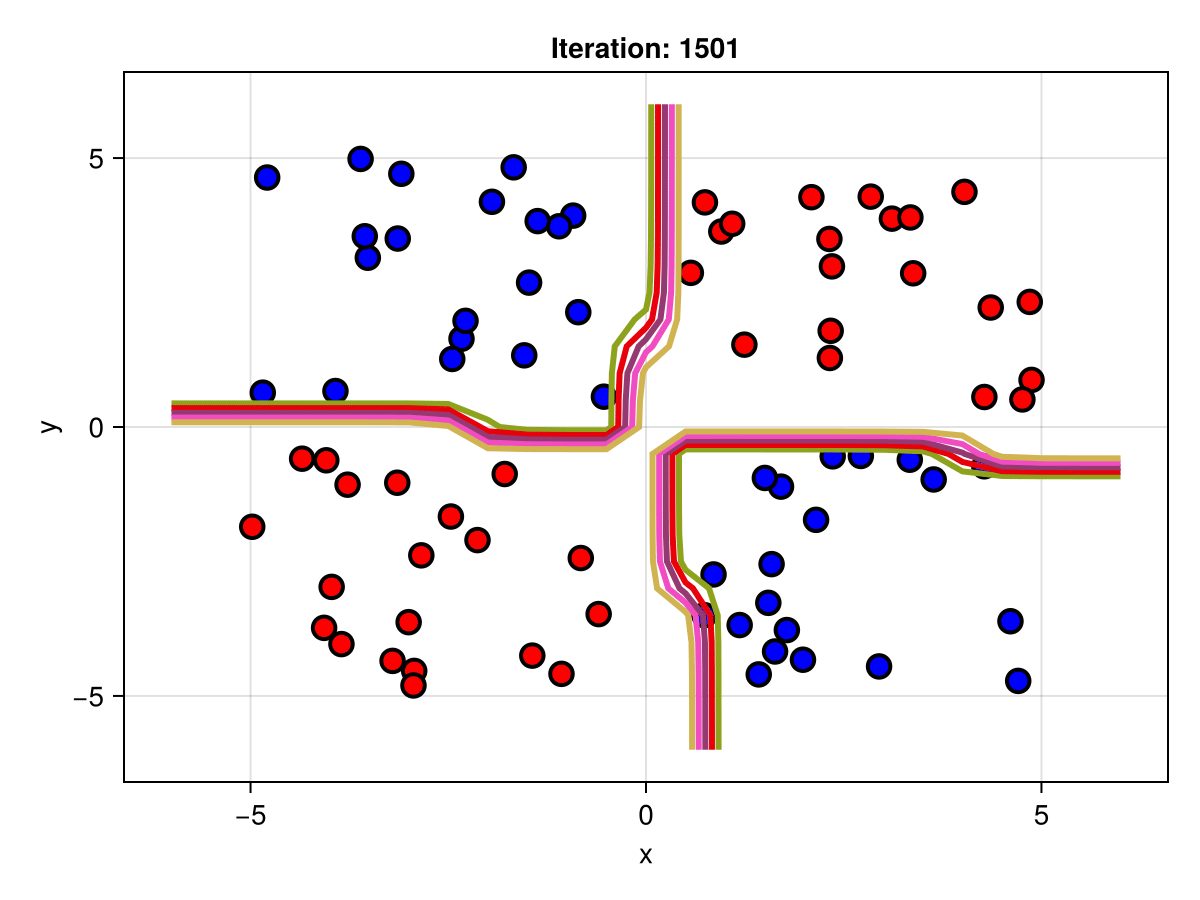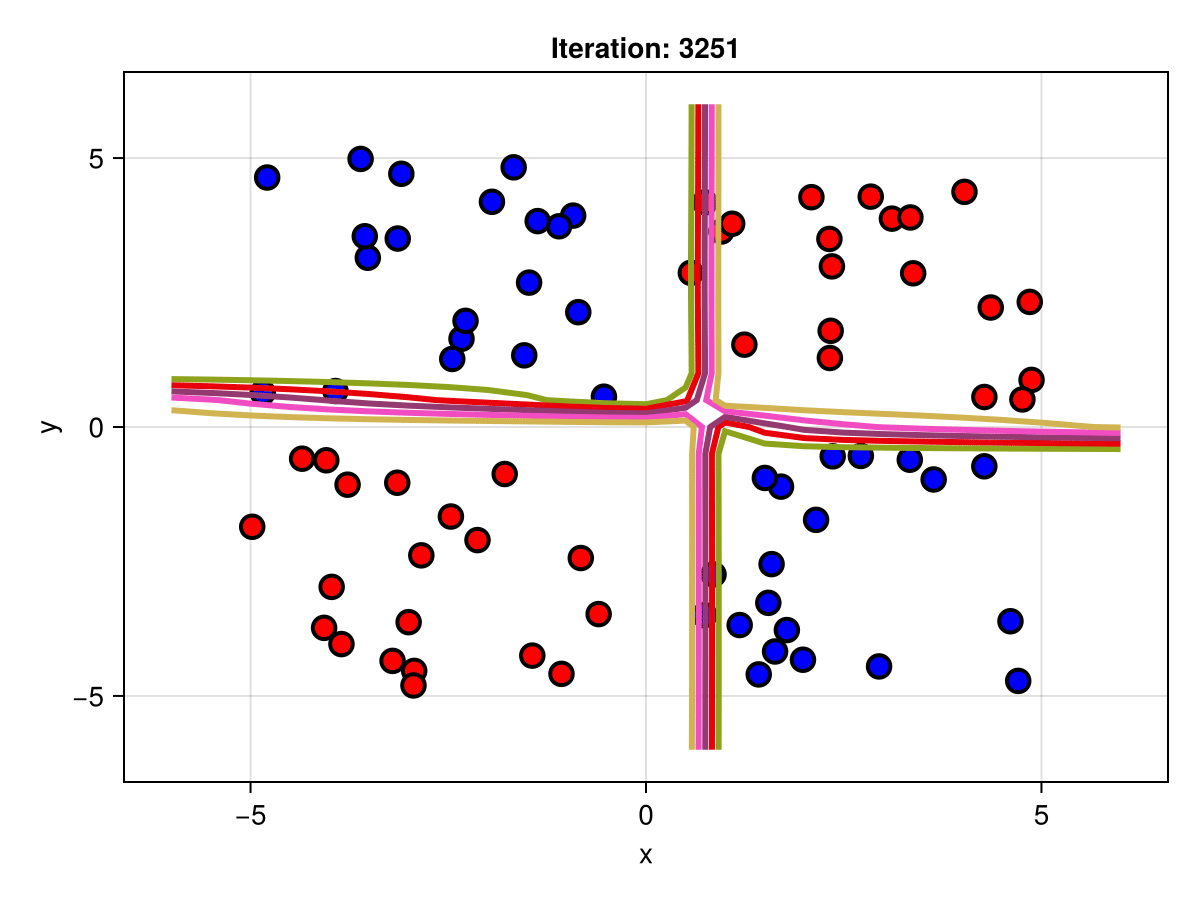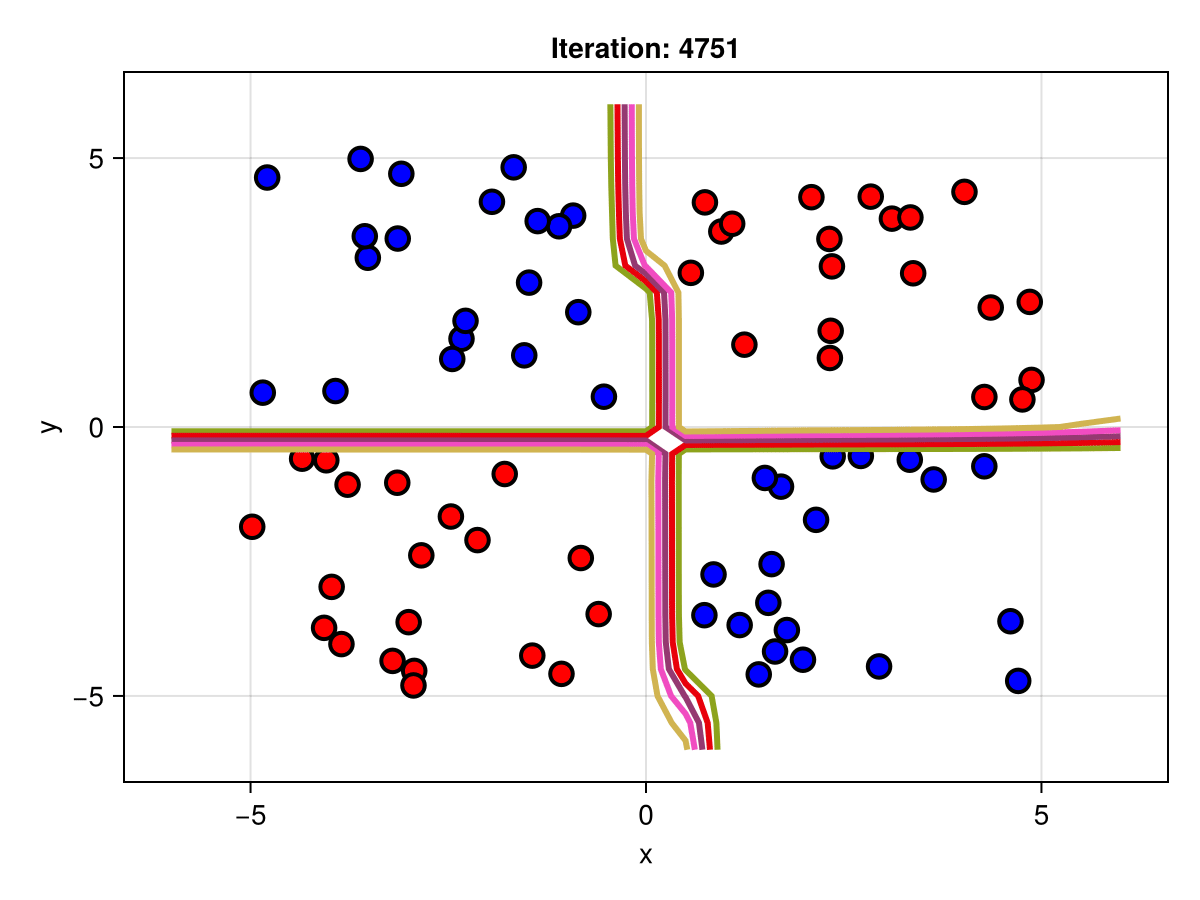
Appendix
using InteractiveUtils
InteractiveUtils.versioninfo()
if @isdefined(MLDataDevices)
if @isdefined(CUDA) && MLDataDevices.functional(CUDADevice)
println()
CUDA.versioninfo()
end
if @isdefined(AMDGPU) && MLDataDevices.functional(AMDGPUDevice)
println()
AMDGPU.versioninfo()
end
endJulia Version 1.11.1
Commit 8f5b7ca12ad (2024-10-16 10:53 UTC)
Build Info:
Official https://julialang.org/ release
Platform Info:
OS: Linux (x86_64-linux-gnu)
CPU: 128 × AMD EPYC 7502 32-Core Processor
WORD_SIZE: 64
LLVM: libLLVM-16.0.6 (ORCJIT, znver2)
Threads: 16 default, 0 interactive, 8 GC (on 16 virtual cores)
Environment:
JULIA_CPU_THREADS = 16
JULIA_DEPOT_PATH = /cache/julia-buildkite-plugin/depots/01872db4-8c79-43af-ab7d-12abac4f24f6
JULIA_PKG_SERVER =
JULIA_NUM_THREADS = 16
JULIA_CUDA_HARD_MEMORY_LIMIT = 100%
JULIA_PKG_PRECOMPILE_AUTO = 0
JULIA_DEBUG = LiterateThis page was generated using Literate.jl.